CodonW is a programme designed to simplify the Multivariate analysis (correspondence analysis) of codon and amino acid usage.. #Multivariate analysis #Correspondence analysis #Acid usage #Codon #Amino #Acid
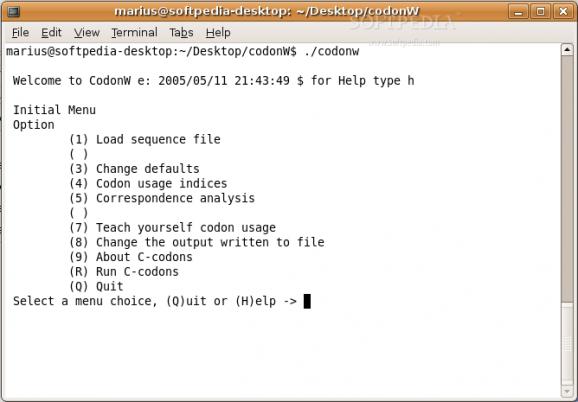
CodonW is a programme designed to simplify the Multivariate analysis (correspondence analysis) of codon and amino acid usage. It also calculates standard indices of codon usage. It has both menu and command-line interfaces.
The CodonW project was written by John Peden in the lab of Paul Sharp, Dept of Genetics, University of Nottingham. John is working in human genetics and is currently employed as ProCardis database manager at the WTCHG in Oxford University.
Here are some key features of "CodonW":
· Written in ANSI compliant C · Menu driven · Extensive number of command line options · Genetic code independent · No limits on
1. number of sequences 2. sequence length
· Calculates the codon usage indices
1. CAI: Codon adaptation index 2. Fop: Frequency of optimal codons 3. Nc: Effective number of codons 4. CBI: Codon Bias Index
· Calculates amino acid indices
1. GRAVY score 2. Aromaticity
· Calculates correspondence analysis of
1. Codon Usage 2. RSCU (Relative synonymous codon usage) 3. Amino acid usage
· Correspondence analysis
1. can include/exclude codons/amino acids 2. can generate detailed reports of trends 3. attempts to identify optimal codons automatically 4. can allow additional data sets to be added 5. records any number of trends
· Calculates gene parameters
1. Gene length 2. GC, GC3s and codon position specific G+C 3. Dinucleotide composition (in all three codon frames) 4. Amino acid usage 5. Relative amino acid usage 6. Codon usage 7. Relative Synonymous codon usage
· Can conceptually translates sequences to protein · Reformats sequence data · Human or machine readable output · Can concatenate genes (whilst preserving reading frame) to calculate
1. overall codon usage 2. GC content 3. Dinucleotide composition
· Generates tables of codon, amino acid, or RSCU usage · Can even help you learn the Genetic code. · and more
CodonW 1.4.4
add to watchlist add to download basket send us an update REPORT- runs on:
- Linux
- filename:
- CodonWSourceCode_1_4_4.tar.gz
- main category:
- Science and Engineering
- developer:
- visit homepage
calibre 7.9.0
Zoom Client 6.0.0.37205
Windows Sandbox Launcher 1.0.0
Bitdefender Antivirus Free 27.0.35.146
paint.net 5.0.13 (5.13.8830.42291)
Microsoft Teams 24060.3102.2733.5911 Home / 1.7.00.7956 Work
ShareX 16.0.1
IrfanView 4.67
4k Video Downloader 1.5.3.0080 Plus / 4.30.0.5655
7-Zip 23.01 / 24.04 Beta
- IrfanView
- 4k Video Downloader
- 7-Zip
- calibre
- Zoom Client
- Windows Sandbox Launcher
- Bitdefender Antivirus Free
- paint.net
- Microsoft Teams
- ShareX